Slow Dynamics and High Variability in Clustered Spiking Networks
Reproduction of Litwin-Kumar & Doiron (2012)
What this is about
Cortical neurons are noisy. They fire irregularly from trial to trial, and their firing rates drift on slow timescales. Standard balanced E/I network models can explain the fast irregularity, but they miss the slow stuff. Litwin-Kumar and Doiron (2012) showed that a simple structural change — clustering the excitatory connections — is enough to get both.
This project is a from-scratch reproduction of their paper. I built a network of 5,000 leaky integrate-and-fire neurons (4,000 excitatory, 1,000 inhibitory) in Brian2, organized the excitatory neurons into 50 clusters with stronger within-cluster connections, and ran the analyses to verify the key claims.
The interesting part
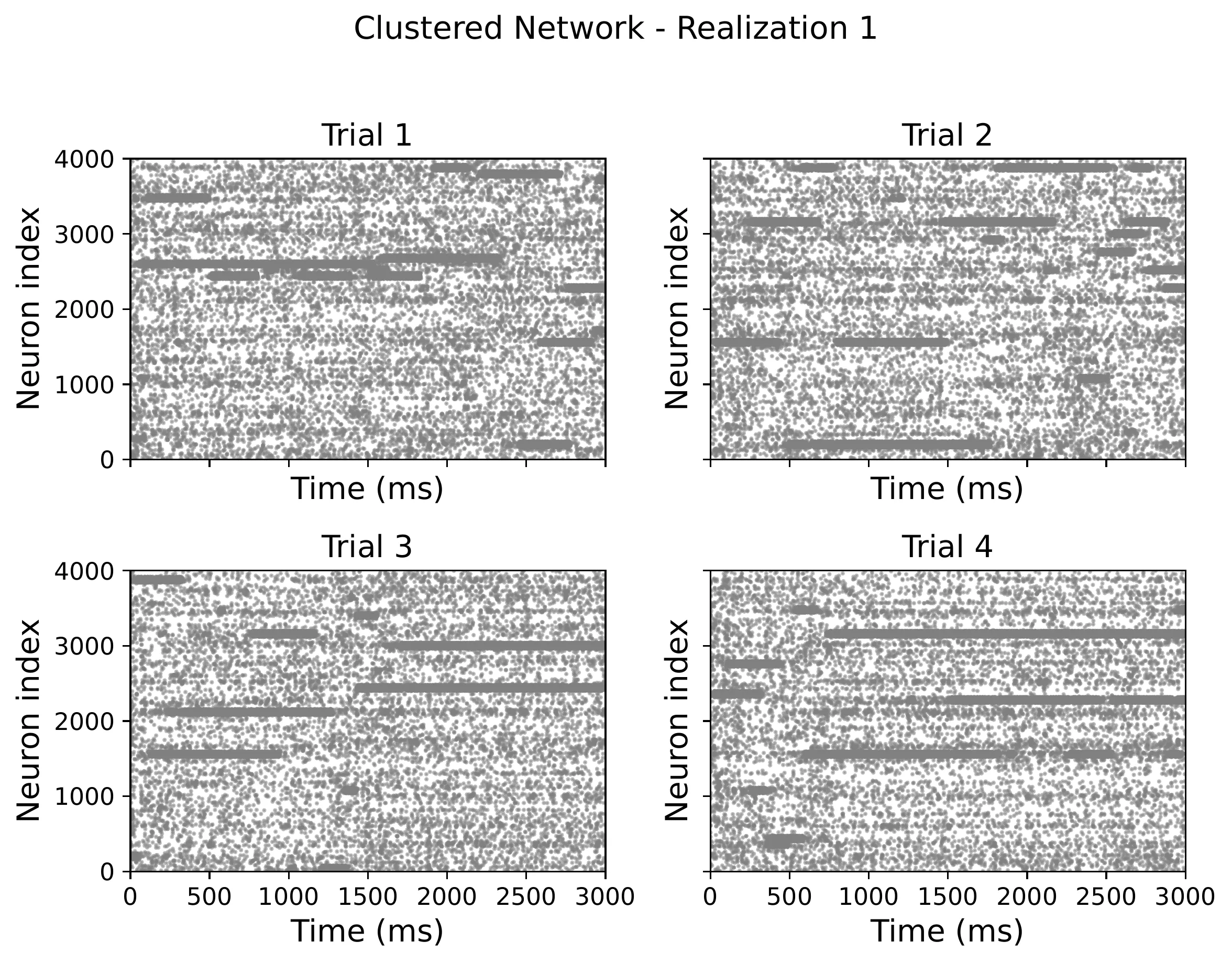
When you strengthen the connections within clusters, the network develops metastable states — clusters spontaneously flip between high and low activity. This looks striking in raster plots: you can see coordinated bursts rippling through groups of neurons.
These metastable transitions produce exactly the statistics you see in real cortical data: Fano factors above 1 that grow with counting window (slow rate fluctuations), weak average correlations but with a heavy positive tail for same-cluster pairs, and — maybe the neatest result — variability that gets quenched when you drive the network with a stimulus, just like in cortex.
There's a sharp transition too. Below a critical clustering ratio, the network behaves like a uniform balanced network. Above it, the metastable dynamics kick in. The paper nails down where that threshold sits.
How it's built
The code is a modular Python package. Simulation runs through Brian2, but the analysis layer is pure NumPy/SciPy — no simulator dependency, which makes it easy to work with the data separately. There's a clean pipeline from parameters to simulation to spike data to statistics to figures.
I tracked five metrics: firing rate distributions, Fano factors, pairwise correlations, auto/cross-covariance timescales, and stimulus response profiles. Everything runs across multiple trials and random seeds for statistical robustness.
Reference
Litwin-Kumar, A., & Doiron, B. (2012). Slow dynamics and high variability in balanced cortical networks with clustered connections. Nature Neuroscience, 15(11), 1498–1505.